Train / Test Splits¶
LAION-fMRI ships with predefined train/test splits so that encoding-model and representation-similarity replications can report comparable generalization scores. The splits implement all three methods of the re:vision generalization framework:
Method 1 — Independent within-distribution. An 80/20 split that covers the image distribution as fully as possible while penalising train/test pairs that are too close in feature space. Use the
tausplit.Method 2 — Out-of-distribution clusters. Five-fold CLIP-feature cluster holdout. Use the
cluster_k5_0…cluster_k5_4splits and average results across folds.Method 3 — Out-of-distribution images. Train on the pool’s regular images, test on the 371 held-out OOD stimuli. Use the
oodsplit. Optionally restrict the test set to specific OOD categories with theood_types=keyword onget_train_test_ids()/get_split_masks().Random baselines (
random_0…random_4) are also bundled — comparing them againsttauquantifies how much of a model’s score depends on train/test similarity.
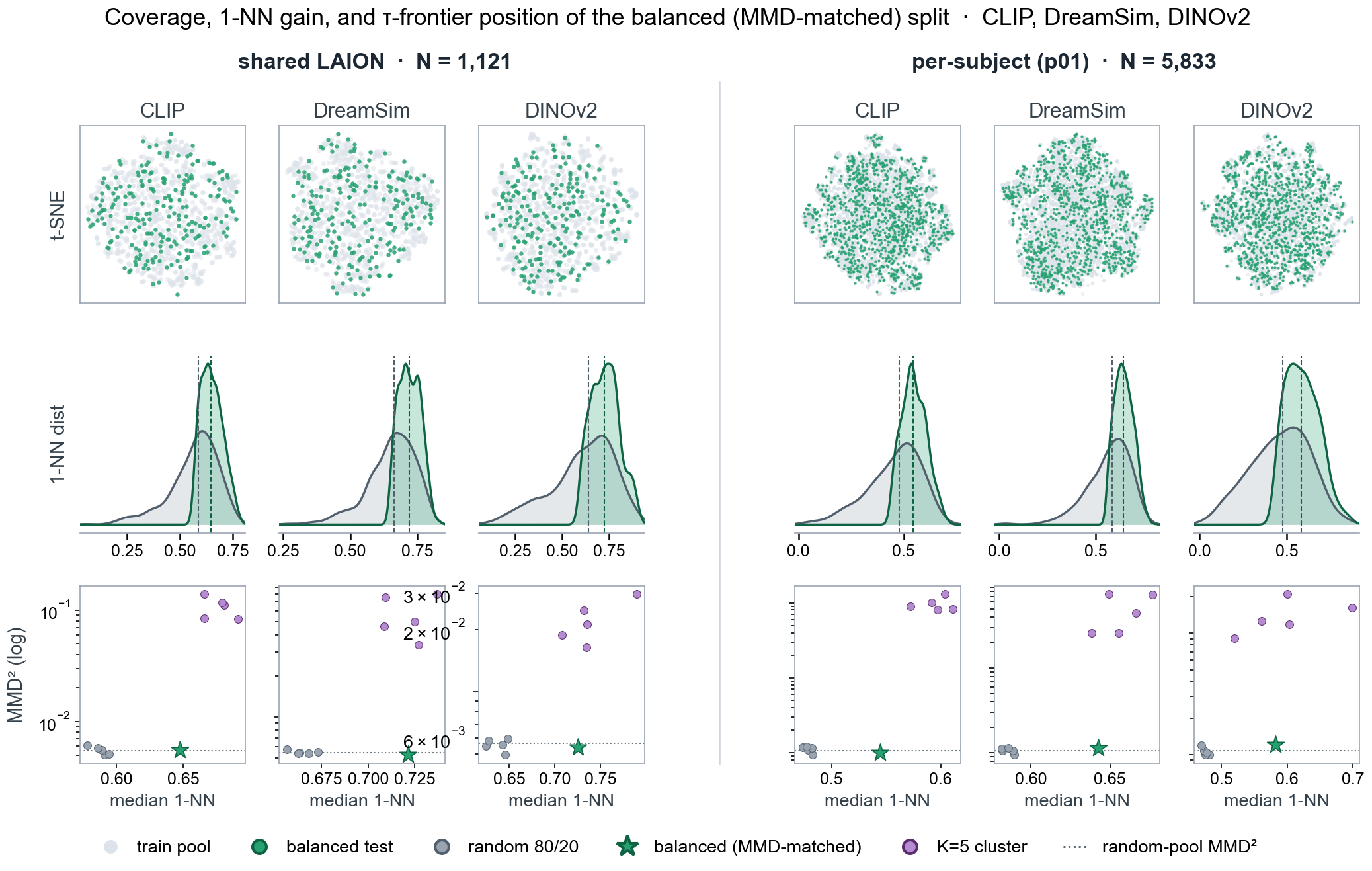
Diagnostics for the bundled tau split, labelled here as the
balanced (MMD-matched) split, in two pools: shared LAION
images (left) and the full sub-01 pool (right). Columns show
CLIP, DreamSim, and DINOv2 feature spaces. The top row shows
t-SNE coverage of the balanced test images (green) against the
train pool (grey). The middle row compares 1-nearest-neighbour
distance distributions for the balanced test split and random
80/20 baselines, with dashed lines marking medians. The bottom
row plots each candidate split by median 1-NN distance and
train/test MMD² (log scale): grey points are random 80/20
baselines, purple points are cluster_k5_* folds, the dotted
line is the random-pool MMD² reference, and the green star marks
the selected balanced split.¶
Pools¶
Every split is bundled for six pools. Pick the one whose
stimulus subset matches the original study — see
list_pools().
"shared"— the 1,121 cross-subject shared images (non-OOD subset of the shared block). Use this when the original study used only stimuli that every subject saw (e.g. NSD- or Conwell-style benchmarks)."sub-01","sub-03","sub-05","sub-06","sub-07"— the per-subject pools (1,121 shared + 4,712 subject-unique images = 5,833 images each). Use a subject’s pool when the original analysis was per-subject and used that subject’s full stimulus set.
Split names¶
Twelve names exist in every pool — see
list_splits().
name |
re:vision method |
notes |
|---|---|---|
|
Method 1 |
The MMD-matched 80/20 nearest-neighbour-distance split. Train and test match at the population level (low MMD), but each test image is kept maximally far from its nearest training neighbour in feature space. Single fixed split per pool. |
|
Method 2 |
Each split holds out one of five CLIP-feature k-means clusters as the test fold. Test sizes vary with cluster population; train + test always sum to the pool. |
|
Method 3 |
Train = the pool’s regular images, test = the 371 held-out OOD shared images. The test set is identical across pools; train varies. See Method 3 — OOD images below for the 9 OOD categories and how to filter to a subset of them. |
|
baseline |
Five seeded uniform-random 80/20 partitions. Use them as a baseline for any generalization metric. |
Split sizes:
random_*andtauare fixed at 80/20 of the pool — that’s 897 / 224 for the shared pool and 4666 / 1167 per subject.cluster_k5_*test sizes vary with cluster population; train + test always equals the pool size.oodtest = 371 for every pool. Train = the pool itself (1,121 forshared, 5,833 for the subject pools).
Loading splits¶
The simplest path is get_train_test_ids(),
which returns the train and test image-id lists. Match those
against the label column of the
get_trial_info() output, then
apply the mask to your betas:
import numpy as np
import pandas as pd
from laion_fmri.subject import load_subject
from laion_fmri.splits import get_train_test_ids
sub = load_subject("sub-01")
# Load betas and trial info as usual (see laion_fmri_package/load).
sessions = sub.get_sessions()
betas_per_ses = sub.get_betas(session=sessions, roi="hlvis")
trials_per_ses = sub.get_trial_info(session=sessions)
# Concatenate across sessions (standard idiom).
betas = np.concatenate(list(betas_per_ses.values()), axis=0)
trials = pd.concat(list(trials_per_ses.values()), ignore_index=True)
# Method 1: within-distribution generalization on the shared pool.
train_ids, test_ids = get_train_test_ids("tau", pool="shared")
train_mask = trials["label"].isin(train_ids).to_numpy()
test_mask = trials["label"].isin(test_ids).to_numpy()
X_train, X_test = betas[train_mask], betas[test_mask]
Matching is by string equality: train_ids / test_ids
are the same strings as the label column entries (e.g.
"shared_12rep_LAION_cluster_1003_i0.jpg"), so no
normalisation is needed.
For convenience, get_split_masks()
collapses those last three lines:
from laion_fmri.splits import get_split_masks
train_mask, test_mask = get_split_masks(trials, "tau", pool="shared")
X_train, X_test = betas[train_mask], betas[test_mask]
Method 2 — five-fold cluster average¶
from laion_fmri.splits import get_split_masks
scores = []
for k in range(5):
train_mask, test_mask = get_split_masks(
trials, f"cluster_k5_{k}", pool="shared",
)
fit_encoding(features[train_mask], betas[train_mask])
scores.append(score(features[test_mask], betas[test_mask]))
m2 = float(np.mean(scores))
Method 3 — OOD images¶
The ood split holds out 371 OOD stimuli as a test set, with
the train side being the pool’s regular images. Every subject saw
all 371 OOD images, so test_ids is the same set regardless of
pool; what changes is what counts as “regular” training data.
from laion_fmri.splits import get_split_masks
# Method 3: train on shared, test on OOD-shared
train_mask, test_mask = get_split_masks(trials, "ood", pool="shared")
fit_encoding(features[train_mask], betas[train_mask])
m3 = score(features[test_mask], betas[test_mask])
OOD categories¶
The 371 OOD images span 9 categories. Discoverable at runtime via
list_ood_types():
type |
n |
what’s in it |
|---|---|---|
|
82 |
abstract shape, colour and digit stimuli |
|
72 |
unusual spatial relations between objects |
|
64 |
natural scenes with unusual content |
|
60 |
zoomed-in / cropped object close-ups |
|
30 |
custom-shot stimuli |
|
24 |
highly saturated / patterned scenes |
|
21 |
textbook visual illusions |
|
10 |
Gabor patches |
|
8 |
natural-image illusions |
Pass ood_types= to
get_train_test_ids() or
get_split_masks() to restrict the test
set to specific categories — useful when only some OOD types
make sense for the original study (e.g. an object-recognition
finding probably shouldn’t be evaluated on Gabor patches):
# Just shape / unusual / cropped (positively biased toward objects)
train_mask, test_mask = get_split_masks(
trials, "ood", pool="shared",
ood_types=["shape", "unusual", "cropped"],
)
# Or one type at a time, to compare per-category generalization:
per_type_scores = {}
for t in list_ood_types():
_, m = get_split_masks(trials, "ood", pool="shared", ood_types=[t])
per_type_scores[t] = score(features[m], betas[m])
Inspection¶
For the full Split object — including
splitter / params / variant metadata — use
load_split():
>>> from laion_fmri.splits import list_pools, list_splits, load_split
>>> list_pools()
['shared', 'sub-01', 'sub-03', 'sub-05', 'sub-06', 'sub-07']
>>> list_splits()
['cluster_k5_0', ..., 'random_0', ..., 'ood', 'tau']
>>> sp = load_split("tau", pool="shared")
>>> sp.n_train, sp.n_test, sp.split_family
(897, 224, 'tau')
>>> sp.params
{'method': 'min_nn_filter + stochastic_mmd_swap', ...}
Split file schema¶
The bundled JSONs live under laion_fmri/splits/data/{pool}/ and
are accessed transparently by the loaders above. Schema, for
reference:
{
"name": "random_0",
"pool": "sub-01 full pool (unique + LAION non-OOD shared)",
"splitter": "min_nn_stochastic",
"params": { "method": "uniform_random", "seed": 0 },
"n_train": 4666,
"n_test": 1167,
"variants": [
{
"variant_id": 0,
"train_ids": ["shared_12rep_LAION_cluster_1003_i0.jpg", "..."],
"test_ids": ["unique_LAION_initial_cluster_2817_i38_p01.jpg", "..."]
}
]
}
See also¶
Stimulus Data — how stimulus filenames map to dataset labels.
GLMsingle Beta Estimates — per-trial beta estimates the splits slice into.