Note
Go to the end to download the full example code.
Quick Start¶
End-to-end walkthrough: initialize, query, download, and load data.
This example touches every step of a typical LAION-fMRI workflow:
Initialize a local data directory
Query what is available in the dataset
Download data for a single subject
Load and inspect the data
For deeper dives into each step, see the focused examples on initialization, querying, and loading.
Initialize the data directory¶
Examples 1, 2, and 4 share one data directory so the licenses accepted here, and the data downloaded below, are reused by the other examples without re-prompting or re-downloading.
import os
from laion_fmri.config import dataset_initialize, get_data_dir
data_dir = os.environ.get(
"LAION_FMRI_EXAMPLE_DATA_DIR",
os.path.join(os.getcwd(), "laion_fmri_quickstart"),
)
os.makedirs(data_dir, exist_ok=True)
dataset_initialize(data_dir)
print(f"Data directory: {get_data_dir()}")
Data directory: /path/to/laion-fmri-data
Query the dataset¶
The bucket is public, so no AWS credentials are needed. Discovery functions read directly from the S3 bucket, so you can see what is available before downloading anything.
from laion_fmri.discovery import describe, get_subjects
print(f"Available subjects: {get_subjects()}")
describe()
Available subjects: ['sub-01', 'sub-03', 'sub-05', 'sub-06', 'sub-07']
LAION-fMRI Dataset
Bucket: s3://laion-fmri
Subjects: 5 (sub-01, sub-03, sub-05, sub-06, sub-07)
ROIs: EBA, FBA, FFA1, FFA2, IPCS, IPS0, LO1, LO2, MPA, MST, MT, OFA, OPA, PPA, SPCS, TO1, TO2, V1d, V1v, V2d, V2v, V3A, V3B, V3d, V3v, VO1, VO2, VWFA1, VWFA2, hV4, laionEVC, laiondorsal, laiongeneral, laionlateral, laionventral, lobjects, mfswords, pSTSfaces, pSTSwords, vobjects
Download one subject – but just one session, in parallel¶
A full subject is several tens of GB. download accepts BIDS
entities (ses, task, space, desc, stat,
suffix, extension) as filters, so you can grab just the
slice you need.
About ses: when set to a session ID, only that session’s
files are pulled – subject-level aggregate maps (the per-subject
noise-ceiling variants, the mean-R^2 summaries, etc.) are
excluded. To pull only those aggregates, use the special value
ses="averages"; combine with a list to pull both. The brain
mask is the one exception – it’s automatically included with any
ses filter, since the loader needs it to mask voxels.
n_jobs runs that many aws s3 cp workers in parallel. The
call is also idempotent – re-running after an interrupted
transfer skips files that already match the bucket size, so you
only fetch what’s missing.
The neuroimaging data and the stimuli are covered by two separate
licenses. On the first download you will be prompted twice –
once for each – and you must type I AGREE each time:
Neuroimaging data (CC0 1.0) – unrestricted use.
Stimuli (closed, research-only) – no redistribution, no commercial or AI/ML-training use.
The acceptances are persisted, so the prompts only appear on the first download into a given data directory.
from laion_fmri.download import download
subject_id = "sub-01"
session_id = "ses-01"
print(f"Downloading {subject_id} / {session_id}")
# Most workflows only need the files the loaders read directly --
# trial info, statmaps, and ROI masks. Asking for the suffix
# subset below keeps a session pull around a few hundred MB
# instead of the multi-GB you'd get pulling everything; drop
# ``suffix`` if you also want the raw GLMsingle model dump or
# the JSON sidecars.
download(
subject=subject_id,
ses=session_id,
suffix=["statmap", "trials", "mask"],
include_stimuli=True,
n_jobs=4,
)
Downloading sub-01 / ses-01
/path/to/laion-fmri/laion_fmri/download.py:217: UserWarning: No objects matching the requested filters under s3://laion-fmri/stimuli/.
fetch_laion_fmri(
Load the subject¶
Once data is on disk, load a Subject
and inspect its sessions and available ROIs. The brain mask is
derived on the fly from the subject-level mean-R^2 file
(..._stat-rsquare_desc-R2mean_statmap.nii.gz) – voxels with
any non-zero GLMsingle fit are considered “in brain”.
from laion_fmri.subject import load_subject
sub = load_subject(subject_id)
print(f"Subject: {sub.subject_id}")
print(f"Sessions: {sub.get_sessions()}")
print(f"Voxels: {sub.get_n_voxels()}")
print(f"ROIs: {sub.get_available_rois()}")
Subject: sub-01
Sessions: ['ses-01']
Voxels: 272080
ROIs: ['EBA', 'FBA', 'FFA1', 'FFA2', 'IPCS', 'IPS0', 'LO1', 'LO2', 'MPA', 'MST', 'MT', 'OFA', 'OPA', 'PPA', 'SPCS', 'TO1', 'TO2', 'V1d', 'V1v', 'V2d', 'V2v', 'V3A', 'V3B', 'V3d', 'V3v', 'VO1', 'VO2', 'VWFA1', 'VWFA2', 'hV4', 'laionEVC', 'laiondorsal', 'laiongeneral', 'laionlateral', 'laionventral', 'lobjects', 'mfswords', 'pSTSfaces', 'pSTSwords', 'vobjects']
Single-trial betas¶
get_betas returns (n_trials, n_voxels) within the brain
mask. For real subjects the brain-mask voxel count is ~270k;
multiplied by ~1000 trials per session, that’s ~1 GB per call.
In practice you should always pass an roi= filter to keep the
array small (face-area ROI, e.g. ~1000 voxels, drops the call to
a few MB).
session = "ses-01"
# Without ROI: heavy but works.
betas_all = sub.get_betas(session=session)
print(f"{session} betas (full mask): {betas_all.shape}")
# Recommended: use an ROI filter.
rois_face = sub.get_available_rois(category="face")
if rois_face:
betas_face = sub.get_betas(session=session, roi="face")
print(f"{session} betas (face ROIs): {betas_face.shape}")
ses-01 betas (full mask): (1044, 272080)
ses-01 betas (face ROIs): (1044, 1100)
Save a derived map back to NIfTI¶
get_betas returns a 1-D (n_trials, n_voxels) array
masked to brain-mask voxels. To round-trip any per-voxel
statistic you compute back into a 3-D NIfTI on disk (so
external tools can read it), pass the array to
Subject.to_nifti: it scatters the values into a
(X, Y, Z) volume and writes the file.
mean_betas = betas_all.mean(axis=0) # (n_voxels,)
mean_path = f"/tmp/{subject_id}_{session}_trial_mean.nii.gz"
sub.to_nifti(mean_betas, mean_path)
print(f"Saved {mean_path}")
Saved /tmp/sub-01_ses-01_trial_mean.nii.gz
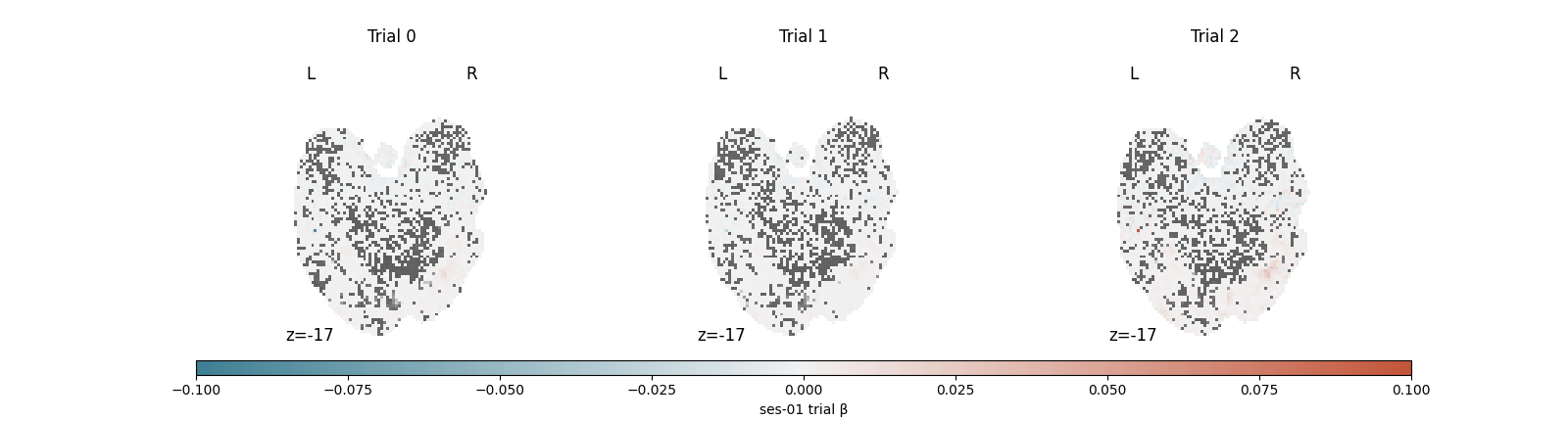
Visualize the first three trials¶
Single-trial betas are inherently noisier than the contrast maps you may be used to: each panel below captures one stimulus presentation rather than an average over many, so don’t expect crisp activation patterns. The view is most useful as a quick sanity check – amplitudes in a reasonable range, signal concentrated in cortex rather than at edges or in white matter, and the three trials looking distinct from each other rather than suspiciously similar.
import warnings
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib.colors import Normalize
from nilearn import plotting
from laion_fmri._paths import r2mean_path
# Nilearn warns about NaN / inf voxels from GLMsingle non-fits;
# they're outside the brain mask and don't affect the rendering.
warnings.filterwarnings(
"ignore",
message="Non-finite values detected",
category=UserWarning,
)
bg_img = str(r2mean_path(get_data_dir(), subject_id))
stat_cmap = sns.diverging_palette(220, 20, as_cmap=True)
trial_paths = []
for i in range(3):
p = f"/tmp/{subject_id}_{session}_trial{i}.nii.gz"
sub.to_nifti(betas_all[i], p)
trial_paths.append(p)
vmax = float(np.percentile(np.abs(betas_all[:3]), 99))
fig = plt.figure(figsize=(16, 4.3))
gs = fig.add_gridspec(2, 1, height_ratios=[1, 0.05], hspace=0.1)
strip_gs = gs[0].subgridspec(1, 3, wspace=0.05)
axes = [fig.add_subplot(strip_gs[0, i]) for i in range(3)]
cbar_ax = fig.add_subplot(gs[1])
for i, ax in enumerate(axes):
plotting.plot_stat_map(
trial_paths[i], bg_img=bg_img, axes=ax,
display_mode="z", cut_coords=[-17],
cmap=stat_cmap, vmax=vmax, colorbar=False,
black_bg=False, threshold=0.5,
)
ax.set_title(f"Trial {i}")
sm = plt.cm.ScalarMappable(
cmap=stat_cmap, norm=Normalize(vmin=-vmax, vmax=vmax),
)
fig.colorbar(
sm, cax=cbar_ax, orientation="horizontal",
label=f"{session} trial β",
)
plt.show()
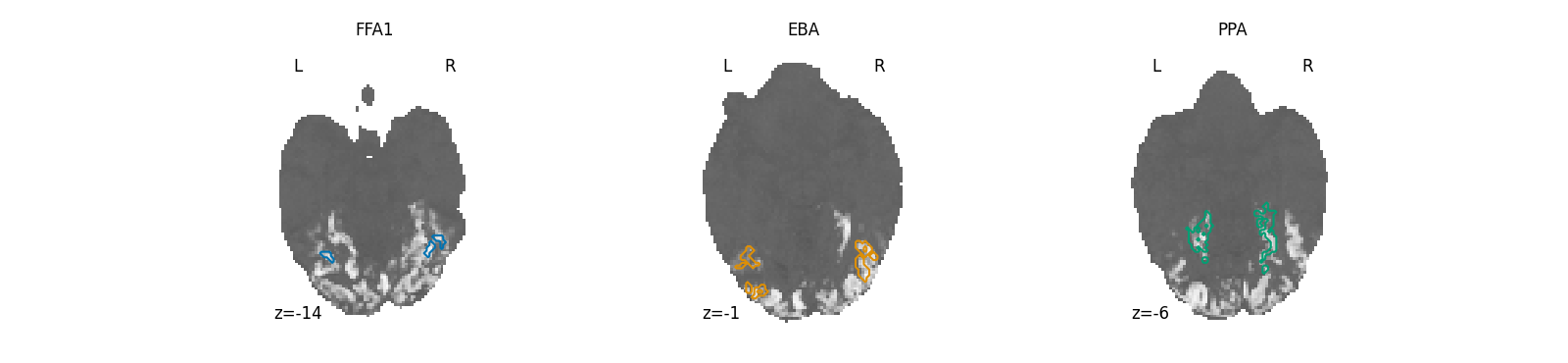
Three category-selective ROIs¶
Face, body, and place ROIs sit in fairly stereotyped parts of ventral temporal cortex, but each subject’s exact localization differs. Glance at this panel to confirm the masks landed where you’d expect – FFA1 in fusiform gyrus, EBA in lateral occipitotemporal cortex, PPA in parahippocampal cortex – before relying on them to filter downstream analyses.
import nibabel as nib
from nilearn.plotting import find_xyz_cut_coords
palette = sns.color_palette("colorblind")
roi_specs = [
("FFA1", palette[0]),
("EBA", palette[1]),
("PPA", palette[2]),
]
fig, axes = plt.subplots(1, 3, figsize=(16, 3.7))
for ax, (roi, color) in zip(axes, roi_specs):
roi_path = f"/tmp/{subject_id}_roi_{roi}.nii.gz"
sub.to_nifti(
sub.get_roi_mask(roi).astype("float32"), roi_path,
)
_, _, z = find_xyz_cut_coords(nib.load(roi_path))
display = plotting.plot_anat(
bg_img, axes=ax,
display_mode="z", cut_coords=[z],
black_bg=False, threshold=0.1, colorbar=False,
)
display.add_contours(
roi_path, levels=[0.5],
colors=[color], linewidths=1.5,
)
ax.set_title(roi)
plt.show()
Per-session noise ceiling¶
NC: shape=(272080,), range=[0.000, 95.076]
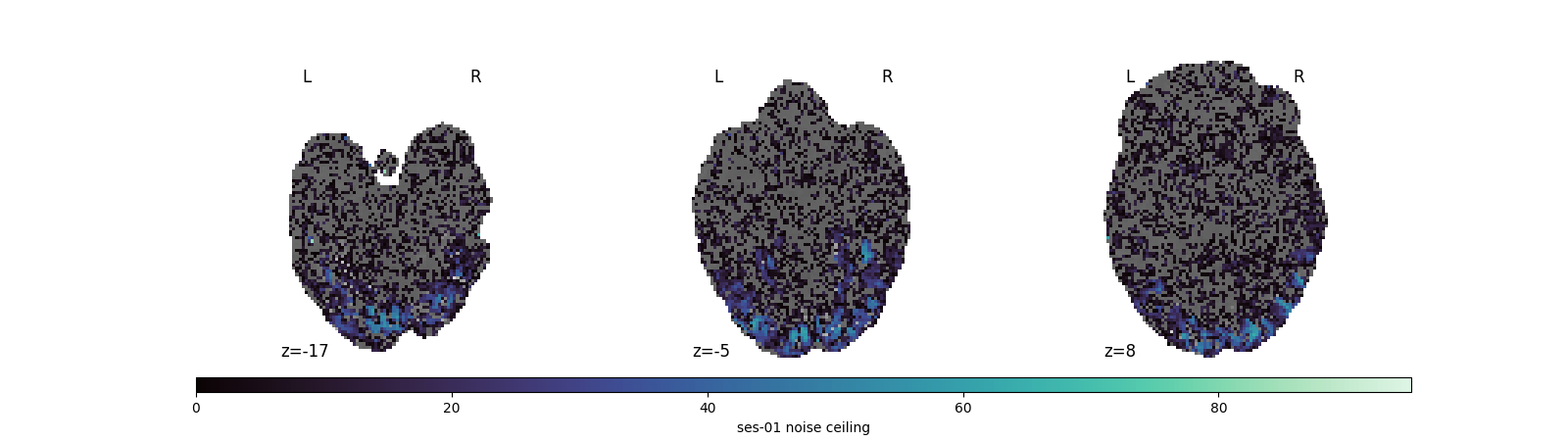
Visualize the noise-ceiling map¶
The noise ceiling tells you which voxels are reliably driven by the stimulus set: high values mark voxels where repeated presentations produce consistent responses, low values mark noise. Looking at this map before any decoding or RSA work helps you decide whether to threshold by NC, restrict to high-NC voxels, or stay with ROI-based analyses.
mako_cmap = sns.color_palette("mako", as_cmap=True)
nc_path = f"/tmp/{subject_id}_{session}_nc.nii.gz"
sub.to_nifti(nc, nc_path)
nc_vmax = float(nc.max())
cuts = [-17, -5, 8]
fig = plt.figure(figsize=(16, 4.5))
gs = fig.add_gridspec(2, 1, height_ratios=[1, 0.05], hspace=0.1)
strip_gs = gs[0].subgridspec(1, 3, wspace=0.05)
axes = [fig.add_subplot(strip_gs[0, i]) for i in range(3)]
cbar_ax = fig.add_subplot(gs[1])
for ax, z in zip(axes, cuts):
plotting.plot_stat_map(
nc_path, bg_img=bg_img, axes=ax,
display_mode="z", cut_coords=[z],
cmap=mako_cmap, vmax=nc_vmax, colorbar=False,
black_bg=False, threshold=0.1,
)
sm = plt.cm.ScalarMappable(
cmap=mako_cmap, norm=Normalize(vmin=0, vmax=nc_vmax),
)
fig.colorbar(
sm, cax=cbar_ax, orientation="horizontal",
label=f"{session} noise ceiling",
)
plt.show()
Stimulus images (when uploaded)¶
Stimuli are forward-compatible: the API is in place but the
images themselves arrive in the bucket later. Until then, the
call below will raise StimuliNotDownloadedError – that’s
the intended signal.
# images = sub.get_images()
# print(f"Images: {len(images)}")
Total running time of the script: (8 minutes 49.078 seconds)